PacBio News
- The new PacBio Sequel chemistry (V3) enables High-Fidelity Long Reads: The chemistry enables significantly longer polymerase reads which can be used to sequence molecules of 10 kb and 15 kb length with 99.9% accuracy. The V2 reagents are discontinued.
- Thanks to the new chemistry the PacBio technology is now optimal for the sequencing of even the longest amplicons. To enable multiplexed amplicon sequencing the DNA Tech Core is now offering two sets of universal barcoded PCR-primers (12×12 and 24×24). These indexes allow multiplexing of up to 576 samples and can be picked up at the lab (for a fee). Please see this FAQ for the protocol, the sequences, and further details: http://dnatech.genomecenter.ucdavis.edu/faqs/how-to-prepare-samples-for-multiplexed-amplicon-sequencing-on-the-pacbio-sequel/
- We have beta-tested V2 of the Express-library prep kit which will be available in early 2019. This kit enables genomic DNA library preparation from as low as 1 ug input. Higher inputs are required for optimal results though. The V2 library prep kit will also improve the performance of genomic libraries (> longer than 10kb) for 20 hour sequencing runs (20 hour movies).
Sequel V3.0 chemistry details:
- The chemistry significantly increases the average lifespan of the polymerase, leading to significantly longer reads and higher yields. Please note that the costs for a V3 SMRT-cell are increased.
- Doubles throughput per SMRT-cell from ~10 Gb to ~20 Gb when (depending on insert sizes)Increases the average polymerase read length from ~20kb to ~30 kb (for inserts > 20 kb)
- Increases the polymerase read length further for shorter insert libraries yielding up to 50 Gb yields and average polymerase read length up to 100kb per SMRT-cell
- Enables high-fidelity long reads by circular-consensus sequencing (CCS) making use of 20 hour sequencing runs and the hairpin adapters: Sequencing 4 passes of library molecule are expected to yield Q20 data and 9 passes Q30 data (99.9% accuracy).
- 20 Hour Movies: Due to the improved polymerase performance it is now suggested to sequence amplicons over 3kb length, Iso-Seq libraries, and of course high-fidelity long read projects (10 to 15kb inserts) with 20 hour movies instead of the standard 10 hour movies. The gains of the doubled run time for longest insert libraries (>30 kb) are marginal at the moment, but will become interesting with the V2 Express-library kit in 2019.
- Please see the information from PacBio and these slides for more details.
These high-fidelity reads can be up to 15 kb long, enabling the detection of all variant types (from SNPs to segmental duplications).
High-fidelity long reads are expected to be preferred for the analysis of metagenomic or population samples, as well as for the assembly of polyploid or highly heterozygous genomes.
The new chemistry significantly improves the quality of the full length RNA-Seq (Iso-Seq) data and amplicon sequencing data.
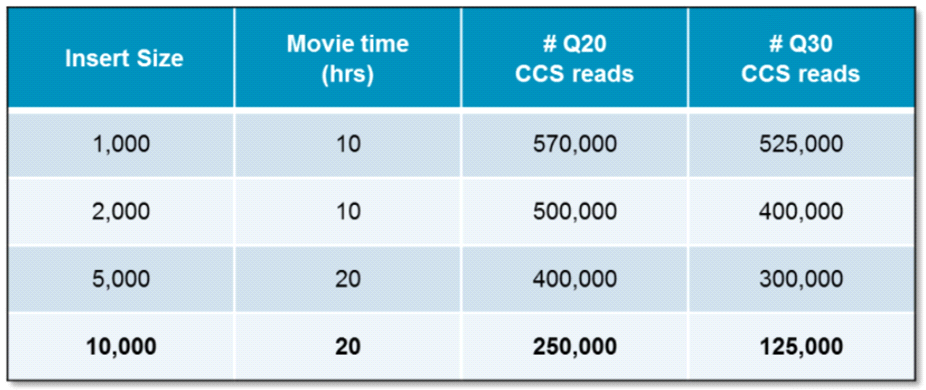
High-fidelity long read quality score data from PacBio (Q20 equals 99% accuracy; Q30 equals 99.9% accuracy):
SMRT-LINK 6.0 software update:
- Structural variant calling now identifies inversions, translocations, and small indels >20bp and allows basepair-precise variant calls
- Iso-Seq3 analysis is more robust and faster
Please subscribe to our Newsletter and follow us on Twitter.