Quick note: The shared equipment is not yet accessible; we offer sample processing on the shared equipment as a service (please contact Siranoosh Ashtari [sashtari@ucdavis.edu] or call the lab).
Please pick a time for a free consultation here
The DNA Technologies and Expression Analysis Core at the Genome Center offers high-throughput sequencing, genotyping, as well as training and consultation. Our goal is to enable access to high throughput genome-wide analyses at economical recharge rates, as a functional extension of your laboratory. We operate on the cost-recovery principle. We employ liquid handling robots to minimize sample handling variation and to provide fast turnaround times. We are a designated Campus Research Core Facility.
We offer all important Next Generation Sequencing (NGS) technologies: Illumina and Element Biosciences short-read sequencing, PacBio and Nanopore long-read sequencing, providing the full spectrum of sequencing options and a wide range of library preparation services for these platforms. Genotyping is performed on the Fluidigm EP1 System for low to medium assay numbers, genotyping-by-sequencing assays, and with a range of genotyping-by-sequencing technologies (e.g. multiplexed amplicon, single-primer extension, reduced representation, and skim-seq protocols). Gene expression analysis is carried out by RNA-seq on short-read sequencers as well as full-length transcript sequencing on long-read systems. Single-cell transcriptome (high-throughput single-cell gene expression profiling) and multi-ome studies are enabled by our 10X Genomics Chromium X system and Parse Biosciences (SPLiT-seq) assays. Spatial transcriptome profiling is supported on the Visium and coming Visum HD platforms with the help of our CytAssist instrument for fresh-frozen, fixed-frozen, and FFPE tissue sections.
We offer annual Illumina and PacBio sequencing library preparation workshops, free consultations on project considerations and experimental design, and custom sequencing library prep solutions. The Core’s equipment (with exception of the sequencers and the genotyping systems) is available to scientists on campus as Shared Equipment for a small usage fee.
We operate PacBio Revio, Oxford Nanopore PromethION, Element Biosciences AVITI, and Illumina NextSeq 500, and MiSeq sequencers, and also sequence on an Illumina NovaSeq X. The sequencers are complemented by a 10X Genomics Chromium X single-cell processor, liquid handling robots (PerkinElmer’s Sciclone NGS G3, two Cybio FeliX, Integra Assist Plus and Viaflo96, Formulatrix Mantis), a Covaris E220 sonicator, and a Fluidigm Access Array System for high-throughput sample and sequencing library preparation. We maintain a LIMS that allows users fast access to sequence data.
Please see our FAQs and information on how to get started with your sequencing and genotyping projects at the Core. Contact us at dnatech@ucdavis.edu with any remaining questions.
Our neighbors at the Bioinformatics Core provide sequence data analysis, statistical evaluations, consulting, and training workshops to help you get the most out of your data. Please contact us for joint consultations with the Bioinformatics Core staff; complete analysis packages, including sequencing and bioinformatics (e.g. differential gene expression, variant calling), are also available.
Our services include:
- MiSeq, and NextSeq sequencing: all run types
- Element Biosciences AVITI sequencing (PE150, PE300, PE80, SE150)
- Illumina NovaSeq X 25B (PE150), 10B (PE150 & SR100)
- DNA sequencing (whole-genome shotgun, targeted, amplicon, exome, ChIP, reduced-representation, custom)
- RNA sequencing (mRNA, total RNA, miRNA, small RNA, 3′ Tag-Seq, custom)
- Methyl-Seq: WGBS (Whole Genome Bisulfite Seq) and RRBS (Reduced Representation Bisulfite Seq)
- PacBio Revio & Sequel II sequencing (whole-genome shotgun long-read, amplicon, and RNA-sequencing)
- Nanopore sequencing on the ONT PromethION
- Gene Expression Profiling via 3′ Tag-Seq — including Differential Gene Expression (DGE) data analysis
- Fluidigm EP1 Genotyping
- Genotyping-By-Sequencing (e.g. multiplexed amplicon, single-primer extension, reduced-representation, skim-seq)
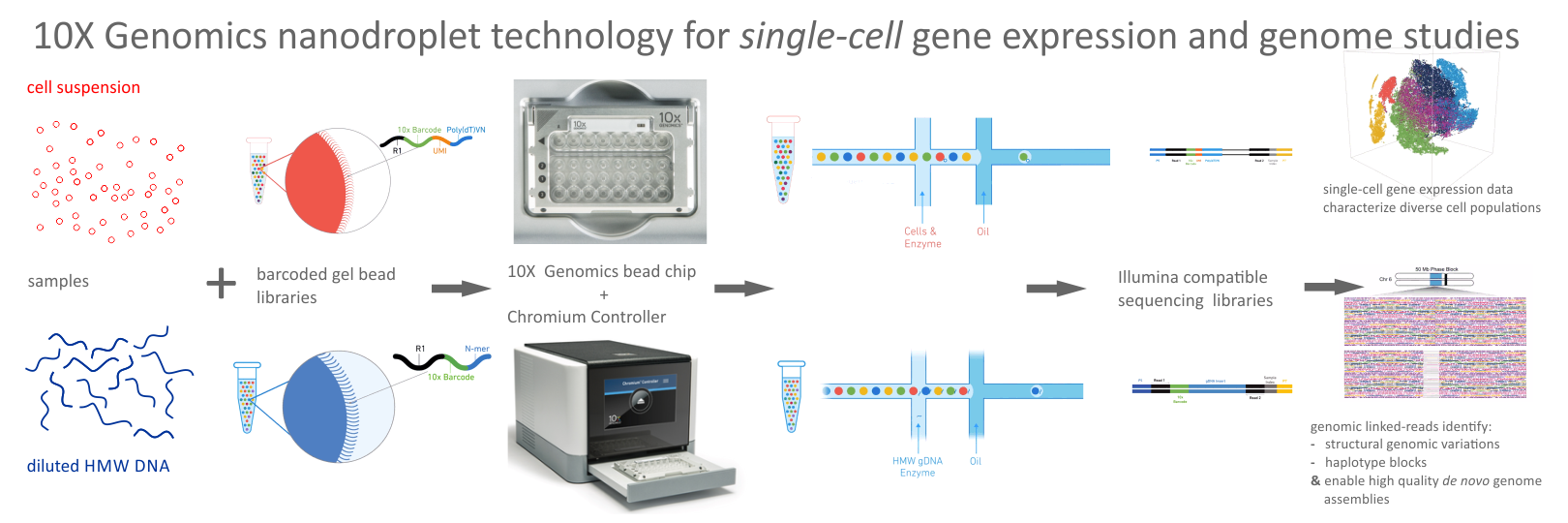
- 10X Genomics Chromium Genome Linked-Read-Sequencing
- Single-Cell Expression Profiling and Multi-Omics
- Spatial Transcriptome Profiling
- Fluidigm Access Array targeted multiplexed amplifications
- High-Molecular-Weight DNA isolations
Sequencing Library Construction and associated services:
- PacBio and Oxford Nanopore long-read DNA sequencing and Iso-Seq RNA sequencing library preparations
- Illumina sequencing library preps, including:
- Genomic DNA library preps
- RNA-Seq library preps (poly-A enrichment or ribo-depletion)
- High-Throughput (HT) library preps (DNA or RNA; starting from 24 libraries)
- miRNA-Seq library preps
- Methyl-seq: WGBS (Whole Genome Bisulfite Seq) and RRBS (Reduced Representation Bisulfite Seq)
- Reduced representation library preps (e.g., genotyping by sequencing)
- Library preps from ChIP samples
- Custom library preparations and sequencing
- Library pooling
- Nucleic acid QC (from single samples to 96-well plates)
- Nucleic acid and library quantification
- Blue Pippin nucleic acid size selection
Please see the Getting Started page to learn how to work with our Core. Please also see our Instrument List. We encourage you to consult our extensive FAQs in case of questions. Illumina has posted NGS tutorials in this “Beginner’s Guide to NGS” .
Our Shared Equipment includes:
- Agilent Bioanalyzer and Tapestation – microcapillary nucleic acid fragment analysis for sample and library QC
- Caliper LabChip GX – a “high-throughput bioanalyzer” (1-384 samples)
- Agilent Femto Pulse – large fragment analyzer using pulsed-field technology, ultra-low input
- Caliper Sciclone G3 – liquid handler robot for high-throughput library preps (up to 96 samples at a time)
- Covaris E220 – high-throughput sonicator (1-96 samples)
- Diagenode Megaruptor – long insert size DNA shearing for PacBio library preps
- BluePippin and PippinHT systems – 50 bp to 50 Kb DNA automated size selection
- Fluidigm Access Array – target amplification for sequencing-ready library prep
- FilterMax F5 Plate Reader – DNA/RNA quantification
- Nanodrop – microvolume DNA/RNA quantification
The Shared Equipment page explains how to get trained.
Acknowledging Our Services
Please support us by acknowledging our services in your publications. We have received NIH funding for the purchase of some of our instruments and this support should be mentioned. Please add a sentence like this to your acknowledgments: “The sequencing was carried by the DNA Technologies and Expression Analysis Core at the UC Davis Genome Center, supported by NIH Shared Instrumentation Grant 1S10OD010786-01.” Acknowledgments such as this constitute big support for future NIH instrumentation grant applications. Please also see our FAQs.
Supporting Your Grant Applications
We support the generation of proof-of-principle data by providing seed grants (please see below) and by offering free-of-charge consultations. We will gladly provide letters of support for your grant application. Please also see our Facilities and Equipment description (Word file) , Facilities and Equipment description (PDF).
Seed Grants
The Genome Center invites proposals for pilot projects that merit support by the Genome Center. These seed grants award up to $2000 per project to spend at any of the Cores; it is also possible to apply for grants at multiple Cores. The Genome Center seed grants are available year-round. They are meant to either introduce UC Davis labs to high-throughput sequencing or genotyping methods, or to help establish new methods. Please see the seed grant FAQ.
Associated Services By Collaborating Cores
- Bioinformatics: The Bioinformatics Core will help you with the analysis of all data generated by our Cores as well as with the statistical interpretation. We highly recommend joint consultations with us and the Bioinformatics Core staff while planning your research project.
- DNA and RNA sample isolation services can be arranged through the Taqman Core.
- 16S Sequencing: We sequence 16S libraries daily. The vast majority of these are customer prepared samples. We do not generate 16S libraries ourselves because UC Davis has a facility specializing in 16S analysis: the Host Microbe Biology Systems Core (HSMBC; also located in our building). The HSMBC offers a complete service ranging from DNA isolation, 16S amplification, sequencing (carried out by the DNA Tech Core), to data analysis. You can also pick selected parts of their service that you require. Please contact the HSMBC manager for details.
- Tilling By Sequencing: The TILLING Core carries out high-throughput mutation screening for rice, wheat, Arabidopsis, and tomato populations.
- Yeast One Hybrid screening: The Yeast One Hybrid Core offers a screening service between promoters and collections of Arabidopsis or maize transcription factors.
- Bionano Saphyr Whole Genome Mapping: The Luo Lab at UC Davis offers whole genome mapping services using the Bionano Saphyr optical mapping technology. This is of special value if the genome assembly for your organism of interest is already of high quality. Please inquire with Ming-Cheng Luo.
- Sanger Sequencing: Since the closure of the DNA Sequencing Facility, Sanger sequencing projects can be submitted to two companies that both have dropoff boxes on campus: Genewiz from Azenta Life Sciences and Quintara Biosciences.
- Cancer Genomics: The Genomics Shared Resource on the Sacramento campus provides general genomics support and custom services for cancer studies.
- Flow Cytometry & Flow Sorting: We work with the Flow Cytometry Core to enable single-cell expression profiling from selected cell populations.
The DNA Technologies and Expression Analysis Core is a UC Davis Campus Research Core Facility. Please subscribe to our newsletter. The general contact address is dnatech@ucavis.edu.