Pacific Biosciences: Realtime Single Molecule Sequencing
 The PacBio Sequel uses proprietary SMRT (Single Molecule Real Time) technology, which enables the real-time detection of nucleotide incorporation events during the elongation of the replicated strand from the non-amplified template. SMRT technology uses nucleotides containing a fluorescent label on the phosphate chain of the nucleotide rather than on the base. Thus, incorporated nucleotides are detected based on the associated fluorophore that is released and dissipated upon cleavage of the phosphate chain, a natural step in the process of DNA synthesis.
The PacBio Sequel uses proprietary SMRT (Single Molecule Real Time) technology, which enables the real-time detection of nucleotide incorporation events during the elongation of the replicated strand from the non-amplified template. SMRT technology uses nucleotides containing a fluorescent label on the phosphate chain of the nucleotide rather than on the base. Thus, incorporated nucleotides are detected based on the associated fluorophore that is released and dissipated upon cleavage of the phosphate chain, a natural step in the process of DNA synthesis.
Real-time detection of nucleotide incorporation events occurs in a nanoscale space called a ZMW (zero mode waveguide), a cavity tens of nanometers in diameter that is fabricated in a 100 nm metal film deposited on a glass substrate. A DNA polymerase, bound to the template DNA strand, is anchored to the bottom glass surface of a ZMW. Laser light traveling through the glass into the ZMW illuminates only the lower 30 nm of it, since the ZMW dimensions are smaller than the wavelength of the light. This allows the selective excitation and identification of light emitted from nucleotides recruited for base elongation that are being held by the polymerase for fractions of a second. 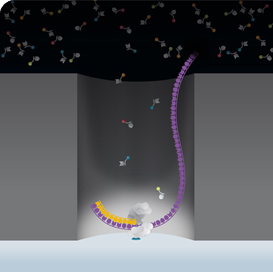 Much shorter signals of unincorporated free-floating nucleotides are recognized as background noise. Each Revio SMRT cell contains an array of 25 million ZMWs, each capable of containing an immobilized strand of library DNA. Collected light pulses from elongating nucleotides at all ZMWs are monitored and analyzed in parallel for the primary analysis (basecalling and quality values). More information describing this instrument and its application is available at these Pacific Biosciences websites here and also here. In contrast to other technologies, it has been shown that the PacBio sequencing chemistry is not sensitive to extreme GC contents. Even long GCC repeat stretches can be sequenced.
Much shorter signals of unincorporated free-floating nucleotides are recognized as background noise. Each Revio SMRT cell contains an array of 25 million ZMWs, each capable of containing an immobilized strand of library DNA. Collected light pulses from elongating nucleotides at all ZMWs are monitored and analyzed in parallel for the primary analysis (basecalling and quality values). More information describing this instrument and its application is available at these Pacific Biosciences websites here and also here. In contrast to other technologies, it has been shown that the PacBio sequencing chemistry is not sensitive to extreme GC contents. Even long GCC repeat stretches can be sequenced.
Getting started info: > Register with the GC > Sample Requirements > Sample Submission
Applications
The DNA Technologies Core at the UC Davis Genome Center operates a PacBio Sequel; our staff has six years of experience in PacBio sequencing. The system currently sequences libraries with insert sizes up to 50 kb. Data produced from contiguous long reads greatly facilitates assemblies and investigation of genomic regions previously difficult to sequence due to long repeats, strong base composition bias, or structural rearrangements. Thanks to the long reads, an unbiased random error profile, and steadily improving bioinformatics PacBio sequencing has become the primary technology for high-quality de novo genome assemblies. Other important applications include isoform sequencing (Iso-Seq; full-length mRNA transcripts — no assembly required), long amplicon sequencing, and bacterial DNA base modification (methylation) studies. Throughput and pricing is competitive, especially for the sequencing of smaller genomes. Pacific Biosciences maintains a site with the most up-to-date citations about the experiments being done with this instrument, and we’re always happy to discuss ideas for employing the PacBio for your project, as well as what’s new and what’s available at our Core.
Services and Performance
We offer complete PacBio library prep, BluePippin size selection and sequencing services. The base calling and secondary analyses like Reads-Of-Insert extraction, HGAP assembly, and IsoSeq analyses of single or a small number of SMRT-cells are included in the service. The Bioinformatics Core is offering in-depth analyses of PacBio sequence data. Samples can be provided as already isolated DNA by the customers. We do offer HMW-DNA isolation services for PacBio sequencing.
Real-time data collection uses a 4 hour movie capture of the SMRT cell. Movies are processed in real time into trace files for each ZMW; base calls and associated quality scores are subsequently determined. These primary analysis files are passed through the secondary analysis pipeline to generate consensus sequence. Each SMRT-cell should generate between 35,000 and 70,000 good (single polymerase) reads. Since the precise quantification of the PacBio libraries before sequencing is not possible, the sequencing of the first SMRT-cell is mainly used to gather titration information and to QC the library. The yield of the first cell thus can be considerably lower than average. PacBio is continually improving read length and quality. The latest chemistry, the V2 chemistry, can produce average reads of up to over 15 kb and longest reads up to 70 kb (for 6 hour movies), given very high quality DNA samples. The official yield estimates provided by PacBio are up to 8 Gb per SMRT-cell (after library titration). We have seen average yields per SMRT-cell of up to 10 Gb (from very good DNA samples). Very long average read lengths are only possible after accurate size selection of the sequencing libraries (as smaller fragments will load preferentially). We offer the automated BluePippin size selection for enriching library fragments longer than 7 or 30 kb.
Please also see our page on PacBio Library Preps and Sample Requirements
Prices
Costs for PacBio sequencing reflect the number of libraries and number of SMRT cells required. Our recharge rates can be viewed here. The listed fees include all labor and reagents. BluePippin size selection is optional (and carries an additional fee), but is highly recommended. Please note our reduced high throughput (HT) recharge rates; these apply if 10 or more SMRT-cells are sequenced per sample or if 6 or more libraries are generated.

